Sequencing The Eastern Yellow Robin: Sex chromosomes with a twist

Painting of the Eastern Yellow Robin by Prof Paul Sunnucks.
Today in Gigascience we published an avian genome assembly with a twist. An Australian team at Monash University discovered unusual, so-called neo-sex chromosomes in the genome of the Eastern Yellow Robin. Being big fans of bird genomes (see our support of the Avian Phylogenomic and B10K projects) it is great to see another one take flight. The surprise finding and the new genome assembly will help the researchers to better understand the genetics of ongoing speciation processes, also involving co-adaption between the mitochondrial and nuclear genomes.
If you are a birder and wish to spot an Eastern Yellow Robin (Eopsaltria australis) you have to travel to Australia, where you’ll find the bird around coastal areas in the east of the continent. Just watch out for a little fellow with a distinctive yellow breast.
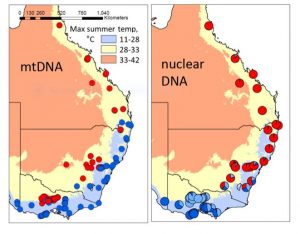
Map illustrating mito-nuclear discordance (adapted from Fig. 1 of Morales et al, 2017).
The Persistence and Adaptation Research Team (PART) at Monash University have chosen the Eastern Yellow Robin as a model and decided to sequence its genome. “Our research group focuses on applying evolutionary concepts to wildlife management to improve the persistence of populations, including how organisms adapt to their environment”, Professor Paul Sunnucks, one of the study’s authors, explains. One major reason why the Eastern Yellow Robin is a particularly interesting system to study evolutionary processes: The species may be in the process of splitting into two, allowing the scientists to get a glimpse of a speciation event as it happens. A smoking gun are peculiar genetic findings that the PART team have discovered over the course of the last couple of years. The previous research showed that there are two genetically distinct lineages of the bird, separated by a mountain range. These two lineages live in different areas with distinct climates: The birds at the coast live in temperate conditions, whereas it is dry inland.
Occasional encounters
Some members of the two lineages do come into contact sometimes and even interbreed. However, depending on which genetic markers the scientists look at, they get a different genetic picture of these encounters, as the PART researchers explain on their project home page:
The mitochondrial genome, which is exclusively inherited by the mother, seems to indicate that there is virtually no gene flow between the two Yellow Robin lineages. However, analyses of nuclear genes reveal that they do mix at least occasionally. The straightforward interpretation is that males and females apparently behave differently in this species.
However, the difference between mitochondrial and nuclear genomes is not so clear-cut, says Dr Alexandra Pavlova:
“A subset of a nuclear genome, mapping to chromosome 1A of Zebra Finch and enriched for genes with functions in the mitochondria, appears to have been co-evolved and co-inherited together with the mitochondrial genome. This makes the Eastern Yellow Robin an emerging model for fascinating evolutionary processes related to evolutionary interactions between the mitochondrial and nuclear genomes.”
The 993 Mb genome assembly that the authors now present in GigaScience (and open data released in our repository GigaDB, here) was achieved by combining Illumina and Nanopore data.
Surprising sex chromosomes
Various analyses, including the anchoring of segments to the published zebra finch genome combined with read depth estimation and k-mer analyses, provided evidence for the presence of neo-sex chromosomes that involve large parts of Chromosome 1A. Professor Sunnucks says:
“We had suspected some sort of chromosome reorganization, but the fact that it turned out to be neo-sex chromosomes was a complete surprise! Neo-sex chromosomes are newly-created or augmented sex chromosomes. They can arise by a range of mechanisms including the emergence of novel sex-determining genes, as well as many different patterns of chromosomal rearrangements, notably fusions between autosomes and sex chromosomes.”
How exactly the previously autosomal regions turned into neo-sex chromosomes is not clear yet, but the authors write in the paper that one relatively simple model is that one copy of chromosome 1A fused with the W chromosome, and the second copy of chromosome 1A became inherited in a neo-Z fashion.
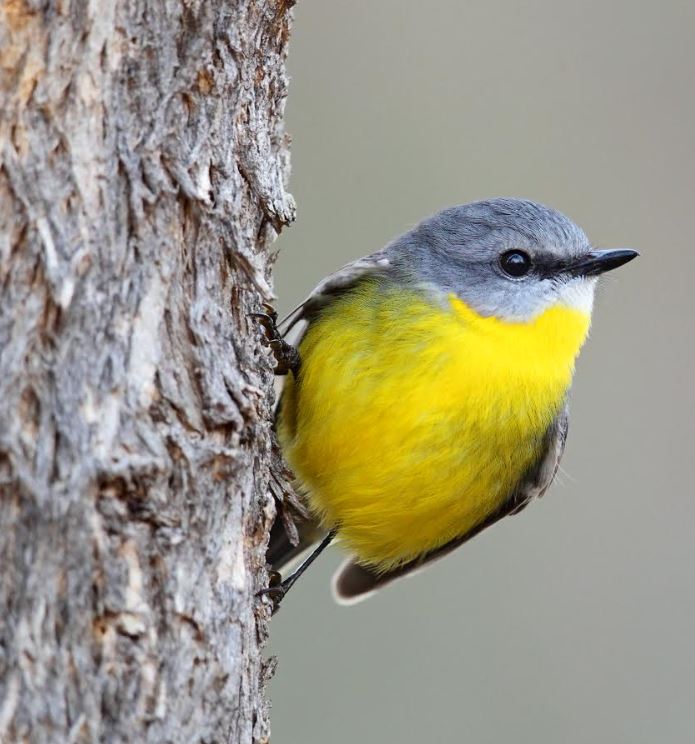

Photo by Chris Tzaros
Prof. Sunnucks puts the findings in context:
“Most sex chromosomes are many tens of millions of years old, by which time it is difficult to observe sex chromosome evolution in action. The neo-sex part of the Eastern Yellow Robin genome is very large and so probably quite recent (we will try to quantify that): it should be subject to ongoing sex-chromosome evolution, including of dispensable genes and some that might be critically important.”
The PART collaboration has come a long way already with this genome project, Alexandra Pavlova recalls:
“The Eastern Yellow Robin project started at Monash University with the publication of the first Eastern Yellow Robin paper in Evolution (Pavlova et al. 2013). In that paper we demonstrated mito-nuclear discordance, ruled out neutral reasons for this pattern and concluded that sex-specific selection must be operating in this species.”
A problem: Most available bird genomes are from males
One of the difficulties the authors had to deal with during this genome project was that the majority of available bird genomes are those of male birds, says first author Dr Han Ming Gan (who is now at Deakin University), adding that the female-specific chromosome is a particularly difficult region to work with:
“Because the W chromosome is not recombining, it rapidly accumulates repetitive material and deleterious mutations, presenting special challenges for genome assembly. “
Another coauthor, Hernán Morales, points out why the neo-sex chromosomes of the Eastern Yellow Robin are quite unique, and how the finding fits into the larger picture of co-adaptive processes in this species:
“Among the few known bird neo-sex chromosomes, the Eastern Yellow Robin one is unique in involving chromosome 1A with its unparalleled concentration of nuclear genes with mitochondrial function. This is particularly alluring because in birds, females are heterogametic as well as the source of mitochondrial DNA, so the female-only neo-sex chromosome will always be coinherited with the mitochondrial genome from mother to daughter to granddaughter and so on, providing for possible intense mitochondrial-nuclear coadaptation.”
As usual, with a genome assembly in hand, the work is not done, but opens the path for new research. Dr Pavlova outlines some of the future plans:
“This includes chromosome-level genome assembly, in-depth work testing whether mitolineages or hybridization impact bird respiration and metabolism, monumental efforts to estimate the fitness in the wild of birds with different genome compositions, and comparative genomics to understand the evolution of neo-sex chromosomes in the Eastern Yellow Robin and its relatives in the Australian Robin family Petroicidae and beyond.”
At GigaScience, we are big fans of bird genomes – for example, during our last ICG prize track, Stefan Prost from the Senckenberg Museum in Frankfurt (Germany) presented a genomic study on five birds-of-paradise species which also revealed interesting evolutionary insights (see Stefan’s talk here). We are excited how genomics contributes to our understanding of bird evolution.
References
Gigascience article:
Han Ming Gan, Stephanie Falk, Hernán E Morales, Christopher M Austin, Paul Sunnucks, Alexandra Pavlova. Genomic evidence of neo-sex chromosomes in the eastern yellow robin, GigaScience 2019. giz111, https://doi.org/10.1093/gigascience/giz111
Supporting data:
Gan HM; Falk S; Morales HE; Austin CM; Sunnucks P; Pavlova A (2019): Supporting data for “Genomic evidence of neo-sex chromosomes in the Eastern Yellow Robin” GigaScience Database. http://dx.doi.org/10.5524/100639
Pavlova, A., J. N. Amos, L. Joseph, K. Loynes, J. J. Austin, J. S. Keogh, G. Stone, J. A. Nicholls, and P. Sunnucks. 2013. Perched at the mito-nuclear crossroads: divergent mitochondrial lineages correlate with environment in the face of ongoing nuclear gene flow in an Australian bird. Evolution 67:3412–3428.
Morales, H., A. Pavlova, L. Joseph, and P. Sunnucks. 2015. Positive and purifying selection in mitochondrial genomes of a bird with mitonuclear discordance. Molecular Ecology 24:2820–2837.
Morales, H., P. Sunnucks, L. Joseph, and A. Pavlova. 2017. Perpendicular axes of differentiation generated by mitochondrial introgression. Molecular Ecology 26:3241–3255.
Morales, H. E., A. Pavlova, P. Sunnucks, R. Major, J. N. Amos, L. Joseph, A. R. Lemmon, J. A. Endler, and K. Delhey. 2017. Neutral and selective drivers of colour evolution in a widespread Australian passerine. Journal of Biogeography 44:522–536.
Sunnucks, P., H. E. Morales, A. M. Lamb, A. Pavlova, and C. Greening. 2017. Integrative approaches for studying mitochondrial and nuclear genome co-evolution in oxidative phosphorylation. Frontiers in Genetics 8:25.
Morales, H. E., A. Pavlova, N. Amos, R. Major, A. Killian, C. Greening, and P. Sunnucks. 2018. Concordant divergence of mitogenomes and a mitonuclear gene cluster in bird lineages inhabiting different climates. Nature Ecology and Evolution 2:1258–1267.
Lamb, A. M., H. M. Gan, C. Greening, L. Joseph, Y. P. Lee, A. Morán-Ordóñez, P. Sunnucks, and A. Pavlova. 2018. Climate-driven mitochondrial selection: a test in Australian songbirds. Molecular Ecology 27:898–918.