Uncovering the tangled roots of plant evolution
 Using big data to understand the tree of life
Using big data to understand the tree of life
New work just published in the Proceedings of the National Academy of Sciences and GigaScience reveals important details about key transitions in the evolution of plant life on our planet, and present a huge cache of computational results, data and tools for plant biologists.
In closing the Origin of Species, Darwin described a “tangled bank, clothed with many plants of many kinds”, and this metaphor is apt for our still tangled understanding of how the key branches of the tree of life fit together, particularly the plant kingdom. From strange and exotic algae, mosses, ferns, trees and flowers growing deep in steamy rainforests to the grains and vegetables we eat and the ornamental plants adorning our homes, all plant life on Earth shares over a billion years of history.
“Our study generated DNA sequences from a vast number of distantly related plants, and we developed new analysis tools to understand their relationships and the timing of key innovations in plant evolution,” said Jim Leebens-Mack, associate professor of plant biology at The University of Georgia and coordinating author of the paper.
As part of the One Thousand Plants (1KP) initiative, the research team is generating millions of gene sequences from plant species sampled from across the green tree of life. By resolving these relationships, the international research team is illuminating the complex processes that allowed ancient water-faring algae to evolve into land plants with adaptations to competition for light, water and soil nutrients. With the aim of generating transcriptome data from over 1,000 species across all the major lineages green plants (although having already exceeded the boundaries suggested by its name, Keith Bradnam points out it could more accurately be called 1.2KP)., this first paper focuses on tools and analyses developed and coming from the first 85 species.
As plants grew and thrived across the plains, valleys and mountains of Earth’s landscape, rapid changes in their structures gave rise to a myriad of new species, and the group’s data also helps scientists better understand the ancestry of the most common plant lineages, including flowering plants and non-flowering cone-bearing plants such pine trees.
“We are using this diverse set of sequences to make many exciting discoveries with implications across the life sciences,” said Gane Ka-Shu Wong, principal investigator for 1KP, professor at the University of Alberta and associate director of BGI-Shenzhen. “For example, new algal proteins identified in our sequence data are being used to investigate how the mammalian brain works.”
“Seeing the impact that 1KP has had inspired us to launch a series of 1000-species projects for organisms like insects, birds and fish, said Yong Zhang, Director of the China National GeneBank (CNGB).
Taming big data
Particularly of interest to us, the project required an extraordinary level of computing power to store and analyze the massive libraries of genetic data, which was provided by the iPlant Collaborative at the University of Arizona, the Texas Advanced Computing Center (TACC), Compute-Calcul Canada, and our collaborators at CNGB, who using the same servers at BGI Hong Kong as us provided early BLAST access to the data.
“This study demonstrates how life scientists are using high performance computing resources to analyze astronomically large datasets to answer fundamental questions that were previously thought to be intractable,” said iPlant’s Naim Matasci.
Computer scientist Tandy Warnow from the University of Illinois Urbana-Champaign and her student Siavash Mirarab developed new methods for analyzing the massive datasets used in the project. “The datasets we were analyzing in this study were too big and too challenging for existing statistical methods to handle, so we developed approaches with better accuracy,” Warnow said.
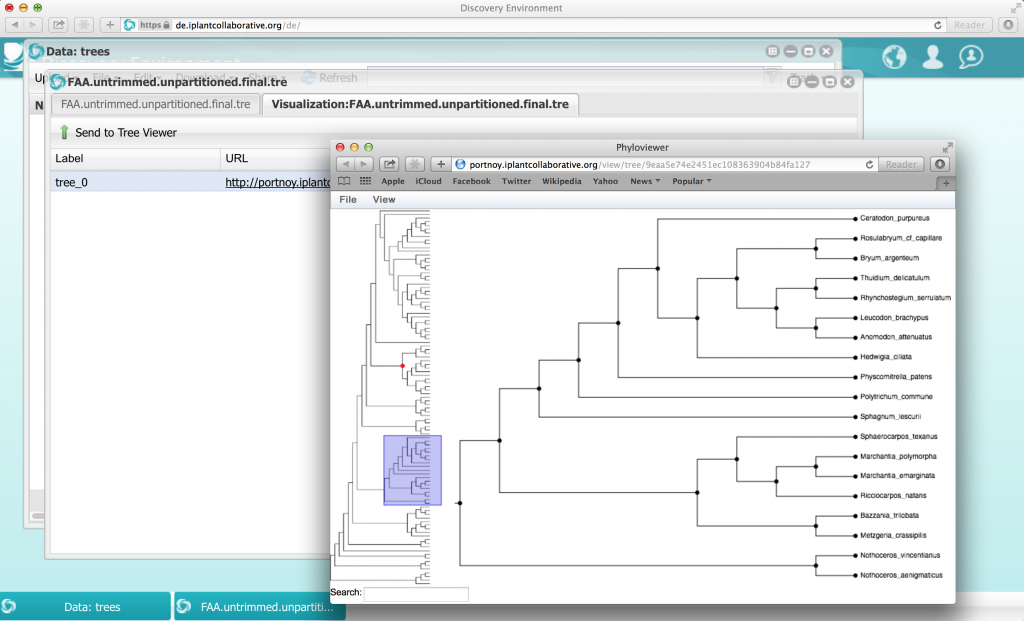
Many organizations, including iPlant, CNGB and the Computational Analysis of Novel Drug Opportunities (CANDO) group at SUNY Buffalo have joined forces to provide web-based open-access to these results. The resources and sequence repositories are described in a companion paper published in GigaScience. The 2009 Toronto Data Release Workshop affirmed and extended the commitments to prepublication release of large genomic data sets which were originally developed in the context of the Human Genome Project. The resulting Toronto Statement encouraged data producers are encouraged to produce a citable statement in which they describe the data set and their intentions in respect of analysis and publication. The GigaScience paper follows in this tradition, linking to the treasure trove of raw and processed data available from the EBI. It also provides more detail on the iPlant Discovery environment, including Phylozoom, a newly developed tree viewer that supports trees with hundreds of thousand leaves and allows for semantic zooming.
Ultimately the researchers hope that their project will not only help us understand the origins and development of plant life, but also provide researchers with a new framework for the study of evolution.
“We hope that this study will help settle some longstanding scientific debates concerning plant relationships, and others will use our data to further elucidate the molecular evolution of plant genes and genomes,” Leebens-Mack said.
Further Reading
1. Wickett et al. (2014). A phylotranscriptomic analysis of the origin and early diversification of land plants. Proceedings of the National Academy of Sciences, U.S.A. Early Access – http://www.pnas.org/content/early/2014/10/28/1323926111.
2. Matasci et al. (2014). Data access for the 1,000 Plants (1KP) project. GigaScience 3:17 doi:10.1186/2047-217X-3-17
Recent comment
Comments are closed.
[…] one of the Darwin finch species among the 48 bird genomes studied, as with the plant 1KP project we recently posted about, this data driven work aims to use the genomics of modern birds to unravel how they emerged and […]