Of Parasites & Protocols, Part 3. Author Q&A with Tony Papenfuss
Following our announcement this month of a new collaboration and integration with protocols.io, we’ve gone into more detail on the first two papers that have utilised this open access repository of scientific methods and collaborative protocol-centered platform. To give some insight into his work, we have one of our author Q&As with Associate Professor Tony Papenfuss, lead author of our scabies genome paper.
Of Mites and Men.
 If you’ve ever seen (or suffered) from a scabies rash, it is in fact caused by thousands of tiny mites, whose very name is derived from the Latin word (scabere) “to scratch”. This makes it a challenging parasite to study, as sample collection ends up with mixed up mess of parasites and the host they’ve been feeding on. Following the recent Q&A with François Olivier Hébert on his tapeworm research, we ask the senior author of our scabies paper similar questions on the difficulties of working with parasites, and experiences being our first authors using protocols.io. Tony Papenfuss heads a lab in the Bioinformatics division at the Walter and Eliza Hall Institute (WEHI) in Melbourne, Australia. Here we ask him about the public health problems of scabies in Australia, and his research trying to produce genomics data to help tackle this.
If you’ve ever seen (or suffered) from a scabies rash, it is in fact caused by thousands of tiny mites, whose very name is derived from the Latin word (scabere) “to scratch”. This makes it a challenging parasite to study, as sample collection ends up with mixed up mess of parasites and the host they’ve been feeding on. Following the recent Q&A with François Olivier Hébert on his tapeworm research, we ask the senior author of our scabies paper similar questions on the difficulties of working with parasites, and experiences being our first authors using protocols.io. Tony Papenfuss heads a lab in the Bioinformatics division at the Walter and Eliza Hall Institute (WEHI) in Melbourne, Australia. Here we ask him about the public health problems of scabies in Australia, and his research trying to produce genomics data to help tackle this.
What are the challenges of working with scabies? How can you collect enough DNA from these tiny parasites to study their genome?
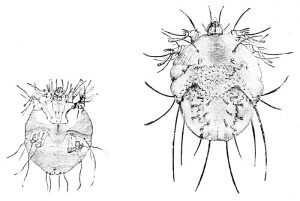 Scabies mites are tiny. Females are less than half a millimetre in length and males half that size again, so we typically sequence samples of about 1000 mites at a time to get enough DNA for next generation sequencing. Their tiny size also means that we cannot remove the gut before sequencing and have to deal with gut flora contaminating the samples.
Scabies mites are tiny. Females are less than half a millimetre in length and males half that size again, so we typically sequence samples of about 1000 mites at a time to get enough DNA for next generation sequencing. Their tiny size also means that we cannot remove the gut before sequencing and have to deal with gut flora contaminating the samples.
Australia is a rich, developed nation ranking in the top 10 countries in the world for their per capita GDP. How big a problem is scabies there, and where did you collect your samples?
Scabies is endemic in many indigenous communities in Australia, but also other disadvantaged communities internationally. In remote communities in northern Australia, up to 25% of adults and 50% of children acquire scabies infections each year, and 7 out of 10 children under 1 year contract scabies. First infestations peak at 2 months of age. Children with scabies do less well, and scabies infections are linked to bacterial skin infections and rheumatic fever. A consequence of this is that indigenous Australians have amongst the highest rates of rheumatic heart disease in the world. This contributes to the significant gap in the life expectancy of indigenous and non-indigenous Australians.
Science is supposedly carried out by standing on the shoulders of giants. How difficult has it been to reuse, adapt and build upon others work in your field?
The challenge with genome assembly in non-model organisms is the iterative exploratory analysis using multiple tools for data cleaning and assembly to construct the best possible assembly, especially in the presence of genetic diversity and contamination by bacterial DNA. It can be a time-consuming and laborious process.
Carrying out an experiment with so many steps involving hosts and parasites, and sensitive pooling, filtering and bioinformatics analysis, how difficult has it been to describing all of this in a textual form in the manuscript?
Writing clear and accurate descriptions of the wet lab and bioinformatics methods is a challenge at the best of times. It is especially hard when the design is complex and iterative exploratory analysis using multiple tools is required. It requires great care and time consuming refinement of the text. I think documenting the methods using protocols.io will make this much easier.
Do you think protocols.io will make this process easier for others to recreate and further build upon your work in the future?
Absolutely.
For more on Tony’s work on the scabies genome project check out a more detailed interview in Australian Life Scientist. See the paper, and the supporting data in GigaDB here. Also check out the protocols below.
References
Mofiz E et al., Genomic resources and draft assemblies of the human and porcine varieties of scabies mites, Sarcoptes scabiei var. hominis and var. suis, GigaScience, Volume 5, Issue 1, December 2016, s13742–016–0129–2, https://doi.org/10.1186/s13742-016-0129-2
Mofiz, E; Holt, D; Seemann, T; Currie, B, J; Fischer, K; Papenfuss, A, T (2016): Draft genome assembly using parasitic mite population NGS DNA sample from mites extracted from host wound environment. Protocols.io. http://dx.doi.org/10.17504/protocols.io.exwbfpe