Taking Spatial Omics into the Next Dimension

A multitude of papers on novel methods for Spatial Omics are published in a cross-journal series launching today in GigaScience and GigaByte Journals.
Spatial Omics is a new field that is taking large-scale data-rich biological and biomedical research into new dimensions. Which is having a significant impact on the fundamental fields of biology and biomedicine. Spatial omics technologies are high-throughput methods for analyzing biological data-based spatial information. It allows researchers to uncover the spatial distribution characteristics in cells, tissues, and organs, which provides a fresh perspective for studying the structure and function of biological systems. This groundbreaking new field has originally centred on spatial transcriptomics, which was named ‘Method of the Year’ by Nature Methods in 2021. For research in this field to continue to progress, new algorithms, methods and tools for spatial omics technology are essential. As well as new data standards and models of sharing this data (see this new commentary covering these challenges). To address these needs, the first articles in a new Spacial Omics thematic series have just been published in GigaScience Press’ open-science journals GigaScience and GigaByte.
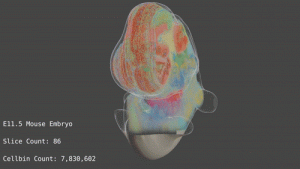
The huge potential of Spatial Omics technology is being held back due to the challenges in handling enormous multi-dimensional datasets. Scientists currently lack available techniques and computational tools for using this novel spatial information even with the ability to re-use existing single-cell data-analysis algorithms. It is therefore crucial to have new customized algorithms and tools specifically created to analyze and interpret spatial omics data. To allow the community to take full advantage of these tools, they must be open-source, easy-to-use, and designed to handle the enormous data volumes produced in these experiments.
To help to establish a place for the community to find multiple new spatial omics analysis methods, GigaScience Press has just published the first batch of articles in a new cross-journal thematic series in our GigaScience and GigaByte journals. These series provide a home for novel spatial omics algorithms, tools, and applications. The openly available pipelines and tools include data preprocessing methods, data quality assessment and improvement, basic analyses, downstream analysis mining, and more. Together, these articles in this ongoing series will help the roll-out and democratize use of this technology by streamlining analyses and providing a toolkit of adaptable open-source tools for others to use and build on. And using our in-house curation experience and data-hosting in GigaDB we can also assist authors to get these complicated works out in an easy and reproducible manner.
One of these just released articles is published in our lead journal, GigaScience, describes a new analysis tool called Siamese Graph Autoencoder (SGAE), an algorithm for detecting spatial domains. SGAE outperforms other methods in terms of capturing spatial patterns and generating high-quality clusters. This enables researchers to resolve anatomical structures such as cortex structures of the brain or gastrulation during mouse embryonic development, with better clarity than other current methods. This groundbreaking new technology pushes the boundaries of what can be studied and discovered. Another GigaScience paper present the imputation algorithm Efficient and Adaptive Gaussian smoothing (EAGS) tool, which improves data quality in highly resolved spatial transcriptomics.
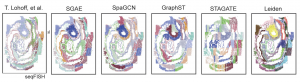
Mouse gastrulation dataset resolved with SGAE and other spatial omics analysis tools.
Articles published in our GigaByte journal address major limiting factors in the adoption of spatial omics research: the availability of workflow systems for data preprocessing. One of articles presents SAW, an already popular (nearly 100 stars on GitHub) tool that processes Stereo-seq data, and which allows better data quality assessment of large spatial transcriptomics datasets. Another paper presents the BatchEval tool, which helps researchers identify and remove batch effects, ensuring reliable and meaningful insights from integrated datasets. In addition to tools for improving data processing, a number of new analysis tool articles are also released in the launch of this new series. These include articles that present the Variable Neighborhood Search (VNS) method, which serves to better cluster cells based on both gene expression and spatial coordinates; and the STCellbin tool, which uses cell nuclei staining images as a bridge to align cell membrane/wall staining images with spatial gene expression maps.
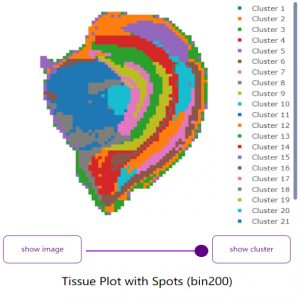
SAW Spatial Omics Analysis workflow, demonstrating spatial clustering for a mouse brain dataset.
This cross-journal thematic series will continue to publish a host of other papers in the coming months, and remains open for submissions of similar open-source, reproducible algorithms, tools and applications. Both GigaScience and Gigabyte aid authors to share due to having in-house data hosting and curation support to provide open-science articles. This encourages others to complement the development of the spatial omics data tool community and continue to promote rapid scientific research progress in this new and growing field.
For more information or pre-submission inquiries, please contact us at editorial@gigasciencejournal.com.
You can read the papers as they continue to be added to the series pages here
GigaByte Journal series page: https://doi.org/10.46471/GIGABYTE_SERIES_0005
GigaScience Journal Series page: https://academic.oup.com/gigascience/pages/spatial-omics-methods-and-applications
References
Cao L et al. Deciphering spatial domains from spatially resolved transcriptomics with Siamese Graph Autoencoder. GigaScience 2024 doi:10.1093/gigascience/giae003
Lv T et al. EAGS: efficient and adaptive Gaussian smoothing applied to high-resolved spatial transcriptomics. GigaScience 2024. doi:10.1093/gigascience/giad097
Gong C et al. SAW: An efficient and accurate data analysis workflow for Stereo-seq spatial transcriptomics. GigaByte 2024 doi:10.46471/gigabyte111
Zhang C. et al. BatchEval Pipeline: Batch Effects Evaluation Workflow for Multi-batch Dataset Joint Analysis. GigaByte 2024 doi:10.46471/gigabyte108
Ivanovic M et al. A Novel Variable Neighborhood Search Approach for Cell Clustering for Spatial Transcriptomics. GigaByte 2024 doi:10.46471/gigabyte109
Kang Q et al. Generating single-cell gene expression profiles for high-resolution spatial transcriptomics based on cell boundary images. GigaByte 2024 doi:10.46471/gigabyte110