Parasite Protocols Part 2. Author Q&A with François Olivier Hébert
 Schistocephalus solidus is as an emblematic study system in parasitology, first discovered by Peter Christian Abildgaard as far back as 1790, as having an extremely complicated life-cycle with multiple developmental states and host species (parasitizing crustaceans, fish and birds). Its fitting that such a classical model system has been used to showcase novel mechanisms of crediting and sharing research protocols in a reproducible manner. Here François Olivier Hébert (Laval University, Quebec) give some insight into his recent Data Note in GigaScience, the methodological challenges of this work, and how easy it was to use protocols.io. For more on this collaboration see also the announcement blogs from us and protocols.io.
Schistocephalus solidus is as an emblematic study system in parasitology, first discovered by Peter Christian Abildgaard as far back as 1790, as having an extremely complicated life-cycle with multiple developmental states and host species (parasitizing crustaceans, fish and birds). Its fitting that such a classical model system has been used to showcase novel mechanisms of crediting and sharing research protocols in a reproducible manner. Here François Olivier Hébert (Laval University, Quebec) give some insight into his recent Data Note in GigaScience, the methodological challenges of this work, and how easy it was to use protocols.io. For more on this collaboration see also the announcement blogs from us and protocols.io.
Schistocephalus solidus has been studied for more than 200 years, so what is still to be discovered about this parasite? Why study the transcriptome?
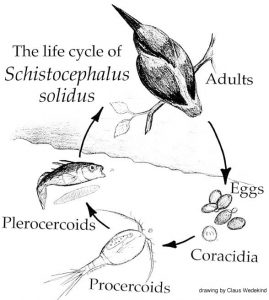 Funny you should ask this question because I am currently reviewing most of the « old papers » that were written on Schistocephalus and I find a lot of parallels between what people found in the 50’s, 60’s and 70’s and what we find with our transcriptomic analysis. In fact, since 1790, people have been extremely interested in the life history traits of this parasite and mainly, its impact on the various hosts is infects. Since then, the scientific community was able to discover various ways to keep the worms in the lab, discover their complete life cycle and develop in vitro techniques that led to important discoveries now widely used in the treatment of various parasite-induced afflictions. So much of the phenotypic and physiological aspects of Schistocephalus have been investigated, but we still don’t know how these biological traits are regulated and managed by the worm. In other words, what are the mechanisms used by the parasite to perform these biological activities? The physiology of the worm allows us to make the link between what happens at the macro scale (e.g. morphology of the life stages) and what happens at the micro scale (e.g. which genes are used at what stage of the infection). So far, we had the morphological and physiological parts, but the micro scale was lacking. One of the approaches to understanding how the functional response of an organism is modulated during its life cycle involves the study of gene regulation. In order to do that, we needed to build a transcriptome and use this reference to investigate how the expression of the genes in the transcriptome is regulated by Schistocephalus. Understanding the link between these complementary levels of biological organization, from the micro all the way up to the macro, will help the scientific community better understand how complex life cycles work in many parasite species, a crucial step towards elucidating how these complex life cycles evolved.
Funny you should ask this question because I am currently reviewing most of the « old papers » that were written on Schistocephalus and I find a lot of parallels between what people found in the 50’s, 60’s and 70’s and what we find with our transcriptomic analysis. In fact, since 1790, people have been extremely interested in the life history traits of this parasite and mainly, its impact on the various hosts is infects. Since then, the scientific community was able to discover various ways to keep the worms in the lab, discover their complete life cycle and develop in vitro techniques that led to important discoveries now widely used in the treatment of various parasite-induced afflictions. So much of the phenotypic and physiological aspects of Schistocephalus have been investigated, but we still don’t know how these biological traits are regulated and managed by the worm. In other words, what are the mechanisms used by the parasite to perform these biological activities? The physiology of the worm allows us to make the link between what happens at the macro scale (e.g. morphology of the life stages) and what happens at the micro scale (e.g. which genes are used at what stage of the infection). So far, we had the morphological and physiological parts, but the micro scale was lacking. One of the approaches to understanding how the functional response of an organism is modulated during its life cycle involves the study of gene regulation. In order to do that, we needed to build a transcriptome and use this reference to investigate how the expression of the genes in the transcriptome is regulated by Schistocephalus. Understanding the link between these complementary levels of biological organization, from the micro all the way up to the macro, will help the scientific community better understand how complex life cycles work in many parasite species, a crucial step towards elucidating how these complex life cycles evolved.
Making your data available in this manner, what do you hope others will do with it?
I hope people download the transcriptome and use it to answer various questions regarding the biology of the parasite. The goal is to use this reference to answer many different questions, whether it is focused on the energy metabolism of the worm, its immunity, reproduction, development, movement, etc. The transcriptome is a very rich and powerful level of biological organization because it contains all of the functional elements used at one point during the life of an organism. I hope people use this reference to study the upper level of biological organization that can be as powerful as the transcriptome: the proteome. Proteins are the molecules that have a direct functional impact in the organism and it all depends on what genes are expressed through the transcriptome and how these gene transcripts are converted into proteins. Proteins are then involved in the functional pathways that give rise to phenotypes, so acquiring some knowledge on what is included in the transcriptome before working on what happens at the proteome and the phenome levels represents a fundamental undertaking. We also hope that people use this reference to perform comparative analyses using many different study systems, which will help better understand the evolutionary aspects of complex life cycles.
How difficult is it to study and collect specimens from all three life cycles? Was one of the parasite/host combinations particularly difficult to work with?
This system is as awesome and fascinating as it is difficult to work with! In order to sample all of the life cycles, we need to make the eggs hatch in the water, then successfully infect enough copepods so that infected copepods can be fed to threespine sticklebacks. We actually have to « force » the sticklebacks to eat one or multiple infected copepods and then we just wait for several weeks and hope that the infection process worked out fine. There are currently no experimental method available to detect the presence of the parasite inside the fish at various stages. The only way to know if a fish is infected is to kill it and dissect it to see if the parasite is located in the body cavity (in between the organs of the host). Unless the parasite has become extremely big over the course of the infection, which tends to create a tremendous abdominal distension in the fish, it is very difficult to tell if it is infected. This very advanced stage characterized by swollen fish bellies is actually the last functional stage of the parasite in the fish, i.e. the infective stage. The problem is that another stage occurs right before that in the fish: the non-infective stage. The truth is, it is super hard to collect Schistocephalus specimens that are at this non-infective stage in the fish because they are very tiny (they only weight a few milligrams) and thus, visually non-detectable when we only look at the general aspects of the host morphology. This non-infective stage was the most difficult stage to work with. Making the eggs hatch, infecting a bunch of copepods, then a bunch of fish and culturing the adult worms in test tubes to stimulate the production of eggs still remain a long process and a tricky protocol to follow because it involves many different steps that rely on complex biological processes and interactions. That is the reason why sometimes, sample sizes are unfortunately quite low.
Collecting good quality RNA for transcriptomics is difficult enough at the best of times. Where there any additional challenges of working in your system?
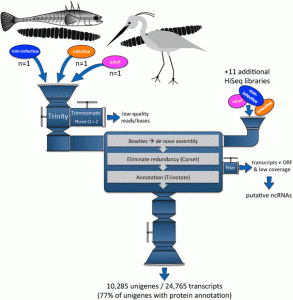 Extracting high quality RNA from these worms was some sort of a challenge because we found something unexpected: cestode RNA does not behave like most « classical » eukaryotic RNA when using standard quality assessment protocols. Very few people have tried to work on the transcriptomics of Schistocephalus and we heard it was quite hard to extract DNA from these worms, probably because of their strong tegument that usually resists the biochemical attack by the digestive enzymes of the hosts. So we thought it would be the same with RNA and we used homemade extraction protocols, only to discover that the RNA profiles used to assess the quality of the samples were systematically « abnormal ». After looking more deeply in the literature, we found that some other people had the same weird profiles and that this was due to a special nucleotide pattern in the sequence of the RNA molecules used to perform the test. This RNA profile can also be found in numerous other cestode and nematode species, as well as in several arthropod species. It took us about four months to realize that, which was kind of a pain because it was supposed to be a standard procedure that usually takes a few days only. All this time we thought it was impossible to extract RNA from these worms, whereas in fact, we had super high quality samples. But that’s just science, it’s all part of the game!
Extracting high quality RNA from these worms was some sort of a challenge because we found something unexpected: cestode RNA does not behave like most « classical » eukaryotic RNA when using standard quality assessment protocols. Very few people have tried to work on the transcriptomics of Schistocephalus and we heard it was quite hard to extract DNA from these worms, probably because of their strong tegument that usually resists the biochemical attack by the digestive enzymes of the hosts. So we thought it would be the same with RNA and we used homemade extraction protocols, only to discover that the RNA profiles used to assess the quality of the samples were systematically « abnormal ». After looking more deeply in the literature, we found that some other people had the same weird profiles and that this was due to a special nucleotide pattern in the sequence of the RNA molecules used to perform the test. This RNA profile can also be found in numerous other cestode and nematode species, as well as in several arthropod species. It took us about four months to realize that, which was kind of a pain because it was supposed to be a standard procedure that usually takes a few days only. All this time we thought it was impossible to extract RNA from these worms, whereas in fact, we had super high quality samples. But that’s just science, it’s all part of the game!
This work is built upon two centuries of research on this species, so how difficult has it been to reuse, adapt and build upon others work?
It is a very interesting process for us to work on such an old and emblematic study system in parasitology because we can benefit from the work of others. When you look at the very old papers on this system, you realize that almost everything has been tried and much have been said on the biology of these organisms. But what is absolutely amazing to understand is that a lot of what have been investigated so far remain to be explained in terms of molecular mechanistic. Basically, the tremendous task that needs to be undertaken consists in bridging what people discovered several decades ago with our molecular perspective and understanding of biological phenomena. This is a golden opportunity to use the scientific method to its full potential because we can read the valuable work of previous scientists and use that information and knowledge to build new research questions that are only accessible through some of the new technologies now widely used in biology (e.g. DNA/RNA sequencing platforms). So we are actively taking part into this extraordinary collective effort of knowledge acquisition. Reusing previous data and results was thus a natural, healthy and essential process to go further, and the real challenge was to read the prolific literature on the subject and identify only a few questions to work with.
Carrying out an experiment with so many steps involving multiple hosts, parasite stages, sample collection, data production and bioinformatics analysis, how difficult has it been to describing all of this in a manuscript?
Describing such a long process of field sampling, experimental infections in the lab using multiple hosts and, of course, the complementary bioinformatic analyses was one of the greatest challenges in this paper. Our main concern is the capacity of anybody reading the paper to reproduce the results that we obtained. From this simple reproducibility concept stems the most convincing aspect of the scientific method: if you can reproduce multiple times the same results, then those results become really strong and from there we can build more solid interpretations. This is why in our paper we emphasized on the detailed description of our methods, as well as all of the scripts and programs used with their corresponding version, including the exact values used for each parameter of the analysis. We were able to achieve that by making all of our homemade scripts, programs and datasets freely available to the public through GigaScience, GigaDB and protocols.io. They represent essential complementary platforms that allowed us to respect our vision of a reproducible science. In order for people to understand why we obtained these results, they have to be able to understand how we obtained them. We do not necessarily wish that people conduct the exact same analyses, with the exact same scripts and the same raw data because that would be replicability, not reproducibility. The important thing here is that people eventually obtain similar results with different methods, different datasets or in different conditions. But in order to compare different methods and their results, we believe it is important to fully explain them. This is why we want to transparently provide to the scientific community with the complete work we did. After all, science is public and belongs to the public.
Do you think protocols.io will make this process easier for others to recreate and further build upon your work in the future?
In this work, we used very precise experimental conditions and sampled biological tissues at very precise moments during the experiment. We think it is crucial for other people to be able to understand what we did and how we did it, so that other research teams can try completely different conditions, slightly change certain parameters or simply improve the protocols and the procedures. Our goal is not that other people reproduce exactly what we did, because there will always be differences between experiments. Whether it comes from differences in the lab equipment that is used or from the populations that are sampled does not matter. What matters is that our methods and protocols serve as a reference point for innovation or improvement and protocols.io offers that possibility. After all, science is an iterative process that inherently generates errors. But, that’s all good, because it is part of the process!
How did you find using the tool, and was the integration of it into our data submission system useful? Are you likely to use it again in the future?
Using protocols.io ended up being quite easy… easier than we thought actually. It requires a little bit more work than not publishing the protocols at all, but in the end, it’s worth it because it involves sharing information and that is a key element to the advancement of science in general. The tool developed by protocols.io was smoothly integrated into the submission process. At first I thought it would be a long and painful procedure, but the platform allows you to enter the information in a familiar format with clear and succinct instructions. I would definitely use this tool in the future.
See Protocols here:
Herbert, F.O.; Grambauer, S.; Barber, I.; Landry, C.R., Aubin-Horth, N. (2016): Protocols for “Reference transcriptome sequence resource for the study of the Cestode Schistocephalus solidus, a threespine stickleback parasite.”. Protocols.io. http://dx.doi.org/10.17504/protocols.io.ew9bfh6
Recent comment
Comments are closed.
Dear FO,
I was really impressed by the relevance of your paper and the angle from which you have decided to approach this problem about the Schistocephalus solidus that has been studied for more than 200 years!!! Understanding the link between the complementary levels of biological organization, from the micro all the way up to the macro, is exactly what needs to be done as this is the approach that is privileged in the study of rare diseases such of the myotonic dystrophy (DM1).
How the functional response of an organism is modulated during its life cycle is precisely the cornerstone of the studies related to the new gene therapies that are at our doorstep.
Your research work is obviously a crucial step towards elucidating how these complex life cycles evolved in the animal world, which may pave the way for the future cure of complex diseases such as cancer and neuromuscular degenerative diseases.
Well-done François-Olivier!
Dr LJ Hébert