Big data, big conferences, big plans: 2019 in review
This is the last blog post of 2019 and it is time again to look back at some of the amazing research published in GigaScience over the past year. Besides handling manuscripts, reviews and data, the editors and curators also attended conferences near and far, they contributed to policy discussions and prepared the launch of a new journal, GigaByte.
More about all these activities below. But first, let’s celebrate some of the science from the journal. As always, it is difficult to select just a few among the many contributions that would be worthy of a special mention, but the following articles certainly stick out:
Sequencing a botanic garden

Sample collection at Ruili botanic garden
We started 2019 with a biggy. Publishing enormous datasets is what GigaScience and its database GigaDB are here for, but the paper and data that scientists from the China National GeneBank, BGI, and the Forestry Bureau of Ruili submitted to us is huge even by our standards: 54 terabytes of sequencing data!
The paper presents genome sequences of 689 vascular plants, representing basically the entire collection of the Ruili botanic garden, thereby tripling the number of plant species with available genome data. Our curators minted 761 individual Digital Object Identifiers (DOIs), making data citation and re-use easy and unambiguous. If you missed it back in January 2019, we presented on it at PAG Asia in June, and you can also read more about the project here.
One thousand plant transcriptomes
 Later in the year, plants again featured big time. The 1KP: One Thousand Plant Transcriptomes Initiative presented their capstone analysis in Nature and accompanied it with a data note in GigaScience, featuring RNA-sequencing data from a whopping 1,173 plant species, and boosting reproducibility by going into more detail on the contamination anaylsis (read the full story here).
Later in the year, plants again featured big time. The 1KP: One Thousand Plant Transcriptomes Initiative presented their capstone analysis in Nature and accompanied it with a data note in GigaScience, featuring RNA-sequencing data from a whopping 1,173 plant species, and boosting reproducibility by going into more detail on the contamination anaylsis (read the full story here).
Birds of paradise

Photo: Tim Laman / naturepl.com
While plants contributed most in terms of data volume to our output in 2019, the animals that populated the journal are hard to overlook either – especially if they are as charismatic and colorful as the birds of paradise which landed in our Data Note section. Coauthor Stefan Prost (Senckenberg Museum, Germany) had presented the work prior to publication at our prize track at the ICG (International Conference on Genomics) in 2018. The article published in January “provides the first glimpse to how genomic evolution is linked to the extraordinary phenotypic variation found in this fascinating group of birds”, co-author Martin Irestedt explained for GigaBlog.

The march of the penguins
Birds – although not flying ones, and clothed in more conservative black-and-white attire – are also the protagonists of another series of data releases. “The Penguin Genome Consortium sequences all living penguin species genomes to understand the evolution of life on the ice”, we announced in a blog post in October. The work is still underway, but the authors provide early access to 19 penguin genomes via GigaDB.
Jellyfish venomes
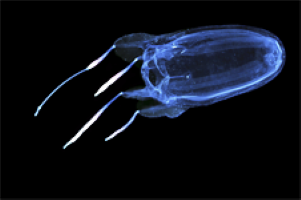 Jellyfish are at least as beautiful as penguins, but some of them are much more dangerous. Joseph Ryan (University of Florida) and his coauthors published genome assemblies of the jellyfish species Alatina alata, Cassiopea xamachana and Calvadosia cruxmelitensis. The analyses are a crucial step to better understand the toxicology of jellyfish venomes (read on in GigaBlog, here, and French readers can read the coverage in Le Figaro):
Jellyfish are at least as beautiful as penguins, but some of them are much more dangerous. Joseph Ryan (University of Florida) and his coauthors published genome assemblies of the jellyfish species Alatina alata, Cassiopea xamachana and Calvadosia cruxmelitensis. The analyses are a crucial step to better understand the toxicology of jellyfish venomes (read on in GigaBlog, here, and French readers can read the coverage in Le Figaro):
A glimpse of prehistoric marine life
In contrast, the marine animals that Mhairi Reid et al. studied can no longer harm anyone. The echinoderms presented in their article died long ago, but they left intriguing fossils that give us a glimpse into long-extinct marine habitats. The researchers at University of Cape Town and Stellenbosch University used micro X-ray computed tomography to get 3D impressions of the fossil-bearing rocks. A great opportunity for our curators to showcase the data using the Sketchfab 3D viewer, integrated directly in the paper. Our data scientist Chris Armit explained:
“3D visualisations are a powerful means of exploring fossil morphology and a Sketchfab 3D viewer, embedded in the paper enables the reconstructed fossiliferous obrution deposit to be explored using a web browser. We previously have been embedding the Sketchfab images in the associated dataset landing pages of our GigaDB database. but this is the first example where Oxford University Press can now embed these in the online version of our paper.”
Mock metagenomes
Papers on microbes also found their way to GigaScience, 2019 prominently in the form of a new “mock metagenome” benchmark, using long read data, from the Nick Loman lab (Birmingham). In an author Q&A (and twitter thread from the first author here) Loman and co-author Sam Nicholls explained that this paper is a starting point to generate reliable trusted data sets, and make them freely available for the community.
A tradition here at GigaBlog is to mark DNA day (April 25) with a special post, this year in the form of a guest blog by Sheri Sanders, a bioinformatician at the National Center for Genome Analysis Support. She dived into the complexities of the decisions one has to make these days when embarking on a sequencing project, using all the amazing tools that are available today (read it here).
Editors and curators on the road again
Apart from publishing research articles, data notes and technical notes, alongside datasets in GigaDB, our editors and data curators also traveled to a wide range of conferences and workshops, with an overarching theme of promoting FAIR principles and open science.

Water cooling the database team (& friends)
One conference we really can’t miss is Intelligent Systems of Molecular Biology) (ISMB). We launched the journal at ISMB 2012 in Boston and tradition requires that we celebrate our birthday during this conference. This year, the event took place in Basel, Switzerland. During breaks, participants dived into the river Rhine to cool down, as central Europe was in the grip of a heatwave. Around 100 delegates, also including several editorial board members, attended our birthday party, where we also revealed our new T-Shirt featuring “Cyber Panda”. Mathematically inspired artist Gregg Helt kindly donated the design.
Woot, woot! @GreggHelt, lookie here! #CyberPanda has been deployed! 🐼 #ISMBECCB #ISMBECCB2019 #ISMB2019.
Oh, we’re gearing up for #BOSC2019 today! 🍐 https://t.co/qvIEimaeY9— Monica Munoz-Torres (@monimunozto) July 24, 2019
- Also during ISMB, we attended the Bioinformatics Open Source Conference (BOSC), another of our favorite events where participants share our dedication to open and FAIR infrastructure.
- Continuing our involvement in the UN’s Citizen Science Global Partnership, our executive editor Scott Edmunds took part as a delegate at a United Nations event in Nairobi – read his report here.
- Our thematic series on Functional Metagenomics (Meta-Func) is still open for submissions and we kept in touch with this community at “Functional Metagenomics 2019” in Trondheim. Read the report of GigaScience editor Nicole Nogoy here.
- Our lead biocurator Chris Hunter is a member of the board of the Genomics Standards Consortium (GSC), which held at meeting in Vienna this year. Read his report here.
- Chris Hunter gave a talk at the 12th International Biocuration Conference in Cambridge – showing how GigaScience and its database GigaDB lead the way for discoverable data (slides here, blog post here).
- Data scientist Chris Armit visited a couple of data-focused events, for example to learn about the latest in super resolution microscopy – an area we covered in some data-rich publications. We would be happy to see more of this in our submissions inbox! Read Chris’s report here.
- Chris Armit also attended a data visualization workshop at EMBL and brought back new insights and fantastic visualization examples.
- In December, Chris also went to ASCB|EMBO 2019, where the cell imaging community also celebrated 100 years since D’Arcy Thompson’s seminal book “On Growth and Form”. Read the report here.
In summary, it was another busy and eventful year for the GigaScience editors and data curators. New things are to come in 2020, including … wait for it …
The launch of a new journal, GigaByte
Together with our partner River Valley Technologies, we already lifted the curtain a bit on this new endeavor (read the press release here, and sign up for updates):
Gigabyte allows extremely rapid publication of short articles focused on non-complex data sets and rapidly evolving research computational tools and technologies. The articles in this journal will be able to be versioned and forked to allow researchers to add new data and information to an article over time without having to completely rewrite the article.
Watch this space for updates! Before we dive into the adventures that 2020 will bring, we wish to wrap up the year by saying
Thank you: To our readers, our authors, our reviewers and the members of our editorial board.
We, and newly crowned (and newly sequenced) New Zealand Bird of the Year hope you continue to support GigaScience and wish you a Happy New Year!
