New beginnings during a pandemic – the COVID year in review

Looking back at this unusual COVID-dominated year, we are happy to report that GigaScience not only kept going under difficult circumstances, publishing outstanding examples of Big Data science and introducing innovations in the review process. We also opened a new chapter with the launch of a sister journal, GigaByte.
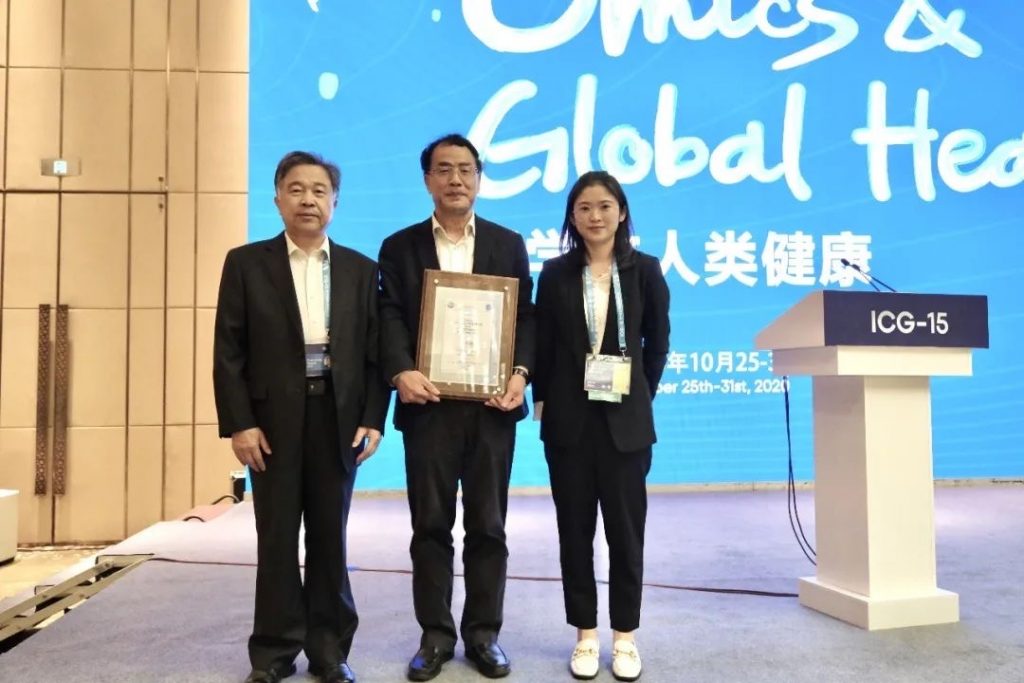
2020 was a year like no other and our end-of-the-year wrap up starts with the one topic that had the world – including the world of science publishing – in its grip: the Coronavirus pandemic. At ICG 15 (the International Conference on Genomics), we recently honored an early hero of this crisis, Prof Zhang Yongzhen, the winner of the 2020 ICG-15 GigaScience Prize for Outstanding Data Sharing during the COVID-19 Pandemic. Dr. Zhang shared the first whole genome sequence of the deadly SarsCoV-2 virus on January 5th, 2020. GigaScience Prize Judge Professor Lachlan Coin (University of Melbourne) explains why this was such an important thing to do:
“Early availability of the genome sequence enabled researchers to start developing vaccines and antiviral therapies even before the virus could be grown in sufficient quantities in cell culture for it to be studied directly.”
Now, as the year comes to an end, first doses of vaccine are routinely administered and a massive roll-out is on the way. Early sharing of data was a crucial step to achieve this milestone. The pandemic is proof that it’s crucial to openly and rapidly share and review research, and GigaScience therefore joined an initiative of a number of journals (the “Rapid Review Initiative”) to streamline and speed up the review process of COVID papers.

Data-centric COVID research
While we are not a specialized infectious disease journal, we also published some data-centric research related to SarsCoV-2. Notably, a scientometric study showing that, in hindsight, research into coronaviruses and other emerging diseases did not get the sustained attention it deserved, compared to other topics such as HIV (read our blog post here). Researching potentially pandemic viruses is a marathon, not a sprint, and the paper served as a warning that scientists and funders need to stay committed to investigating the threat of emerging pathogens.
Another important COVID-related publication was a Technical Note, reporting on an open source, cloud-based tool called IDseq . The platform helps to rapidly detect, identify, and track emerging pathogens such as SARS-CoV-2. “IDseq can be thought of as an early warning radar for emerging or novel infectious agents,” said Joe DeRisi, Co-President of the Chan Zuckerberg Biohub whose research lab at the University of California, San Francisco initiated the IDseq platform. Scientists in Cambodia used the tool to confirm and sequence the whole genome of the country’s first case of COVID.
New beginnings
Thankfully however, 2020 was not all about disease, lockdowns and cancelled travel plans. On a much more positive note, 2020 was also the year we launched our newest baby, GigaScience’s sister journal GigaByte. The new journal is a home for short, focused, data-driven articles. GigaByte is of course an open-access, open data journal and enables rapid publication using new custom-built, end-to-end publishing technology.
“End-to-end” here means that the entire process from submission, review, data curation and production is handled through one streamlined pipeline, making it possible to publish at the speed of research. To make this happen, we worked with our partner River Valley Technologies to deploy a completely new submission and publication platform, also integrating data curation steps. This allows nearly immediate online publication on acceptance, as well as functionality to update published articles.
In the process of launching the new journal, we also changed our organizational structure, giving birth to GigaScience Press as a fully-fledged publisher.
GigaByte waives APCs for the next three months
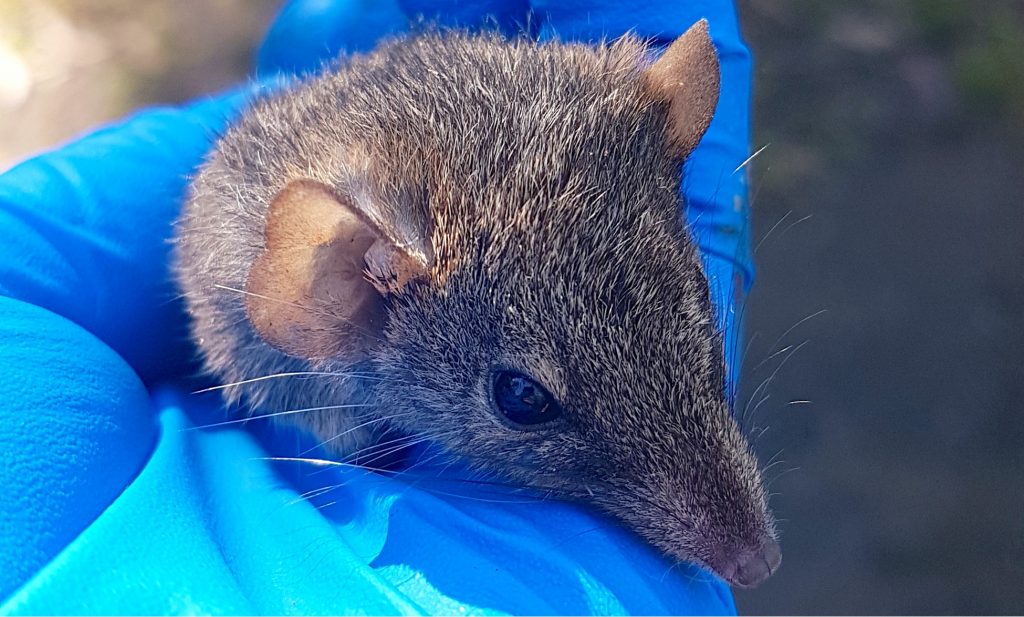
Like GigaScience, our new journal will be a platform to innovate and bring in new features – watch this space for updates! We are happy that the new journal already got a good number of submissions and started publishing now. Among its first articles is genomic work on Antechinus, a genus of small, mouse-like marsupials – see also the Q&A and video abstract with the author).

If you want to become a GigaByte author, there’s good news: For the next three months we are waiving article processing charges for all new GigaByte submissions.
A digital birthday party and looking back at our first eight year
At the end of the year we usually also love to recollect our travels to conferences and fondly recall face-to-face meetings with editorial board members, reviewers and authors. Well, not much travel was possible this year, but we still reached out to various communities thanks to the blessings of virtual conferences.
Even our traditional birthday party had to go digital – in the form of a video (below) and #gigascienceat8 social media lookback over our first 8 years.
Also virtually, our data scientist Chris Armit attended the Human Cell Atlas COVID-19 Virtual Symposium, learning about the mechanisms of COVID-19 disease.
Keeping us updated on the latest developments in the genomics field, and hearing some of our editorial board members speak, we attended at Biodiversity Genomics 2020 (also presenting there on GigaByte) and (back in the mists of time when you could attend in-person meetings) the straightforwardly-named Plant and Animal Genome Conference
Code: checked
While a lot of our attention with respect to the mechanics of peer review was devoted to launching GigaByte, we also introduced some innovations for its established sister GigaScience. Notably, we started to work with the CODECHECK initiative to publish software-heavy articles with certified reproducibility. In short, here’s how it works: After the “code checker” assesses a piece of software, independently time-stamped runs are awarded a “certificate of reproducible computation”, which we display alongside the published article. Our first example was a Technical Note on a machine learning tool called ShinyLearner”. You can check out the paper here and the CODECHECK certificate here.
Another innovative first for us– rather technical in nature, but still important – was the way we handled the assessment of controlled access data during the review of an article by Matthieu Foll et al. . As a journal focused on reproducibility of research, GigaScience has a strict open-science policy. Peer reviewers need access to the data and software supporting our papers. Can this work if sensitive human data is not publicly available to everyone, for ethical and consent reasons ? The answer is yes – because a Data Access Committee (DAC) allowed qualified peer reviewers’ access to inspect the data.
2020’s featured animals: Esperanza, TJ Tabasco and the giant squid
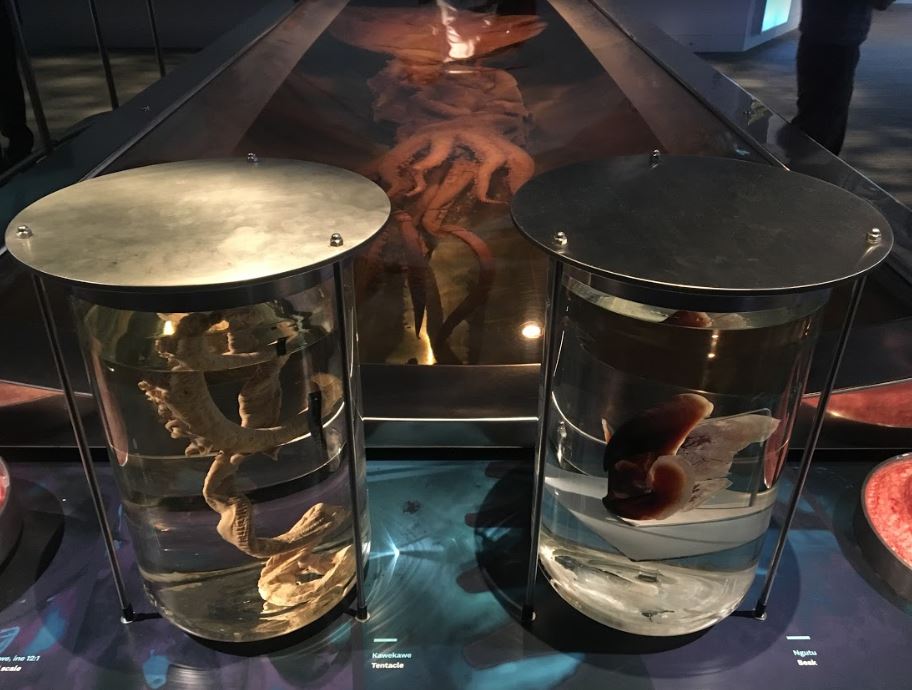
No end-of-the year post can be complete without highlighting at least some of the amazing research and data we published in 2020. We started the year big – rather, gigantic – with a report on the genome of the giant squid Architeuthis dux. A ten-armed invertebrate that is believed to grow up to 13 meters and weigh over 900 kg – no wonder the media loved this, with coverage all over the globe.

For DNA day 2020 we blogged about two special cattle papers (here and here): Rice et al. showed that “trio binning” – that is, using genetic data of a “trio” of two genetically diverse parents and their F1 offspring – can massively improve the assembly process. The authors used a hybrid between cattle and yak (a Yakow? a Yattle?), with the genetic difference between them helping in a clever assembly strategy. Along similar lines, we also published a new cattle reference genome, going back to the genetic material of a famous cow called “Esperanza”, which was also used to produce the first cattle genome back in 2009.
Other species that got new, higher quality versions of established reference genomes include pig (featuring TJ Tabasco, which was the first pig to have its genome sequenced) and the German Shepherd dog.
Mexican caves are at the origin of another research highlight of 2020: A group of Mexican scientists sequenced and assembled the genome of an adult male Lesser Long-nosed Bat (Leptonycteris yerbabuenae) at high quality. They then got sequence data for four more bat species with different feeding habits and detected gene family expansions and contractions that have functions related to different types of diet.
If you wish to learn more about the importance of bat genomics, don’t miss the TED talk of our new Editorial board member Emma Teeling (University College Dublin).
It’s Christmas Jim … but not as we know it.
With the new year around the corner, what the future will bring? Star-Trek-style tricorders maybe. Ok, the kind of handheld universal diagnostic tool used by Dr McCoy may still be science fiction for a long time. But who would have thought that it will be possible in 2020 to sequence entire genomes in a device that’s hardly bigger than a USB stick? And now you can even analyse the results on the go in your smartphone, by using the iGenomics application developed by Aspyn Palatnick, who started to work on the project as a high school student with guidance from our editorial board member Mike Schatz (see our blog post and the authors’ video below).
2020 was a difficult year for most of us, not least for scientists whose projects were held up by lockdowns and other virus-related disruptions. This year, we are especially thankful for all our amazing supporters – first of all our reviewers and members of our editorial board – who contributed their expertise and advice in these difficult times.
Thank you: To our readers, our authors, our reviewers and the members of our editorial board. We wish you all a happy and healthy 2021.
